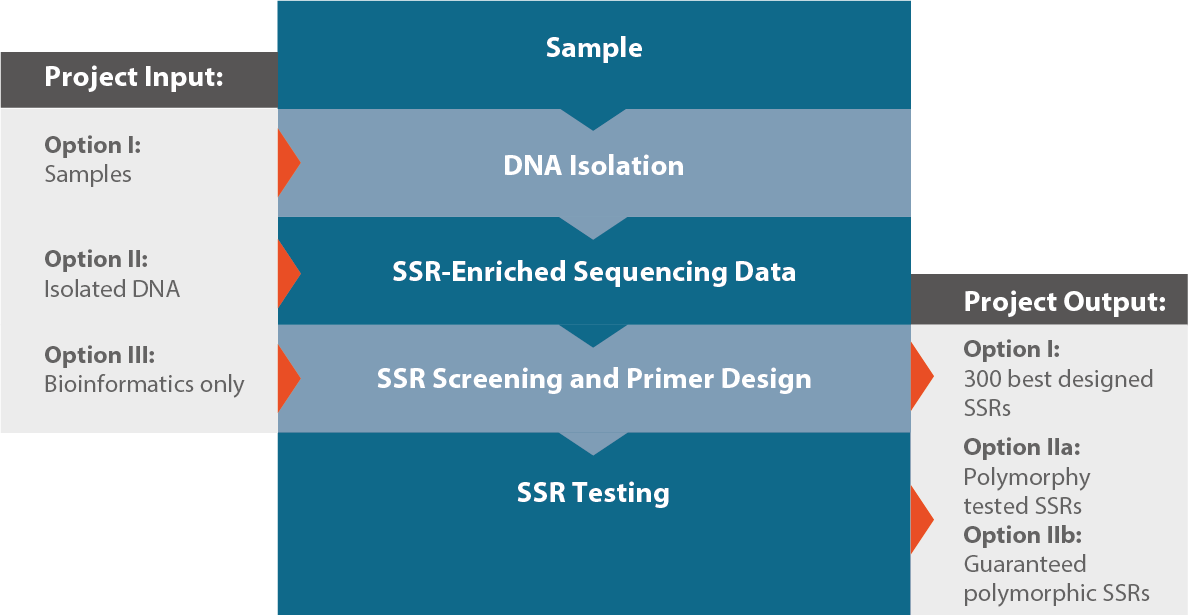
Frontiers | Development of a Large Gene-Associated SSR Marker Set and in-Depth Genetic Characterization in Scarlet Sage
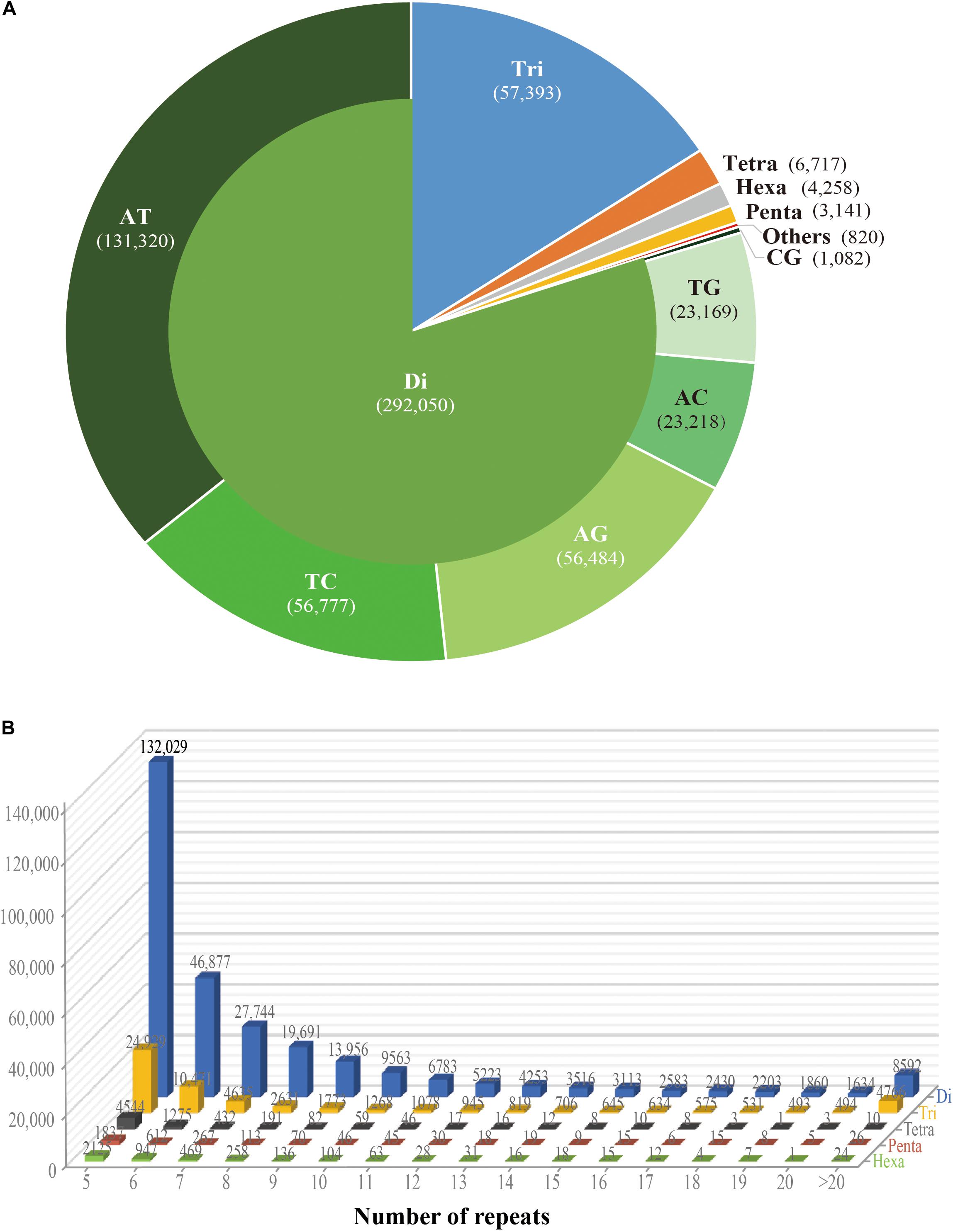
![Development of genic-SSR markers by deep transcriptome sequencing in pigeonpea [Cajanus cajan (L.) Millspaugh] | BMC Plant Biology | Full Text Development of genic-SSR markers by deep transcriptome sequencing in pigeonpea [Cajanus cajan (L.) Millspaugh] | BMC Plant Biology | Full Text](https://media.springernature.com/lw685/springer-static/image/art%3A10.1186%2F1471-2229-11-17/MediaObjects/12870_2010_Article_802_Fig1_HTML.jpg)
Development of genic-SSR markers by deep transcriptome sequencing in pigeonpea [Cajanus cajan (L.) Millspaugh] | BMC Plant Biology | Full Text
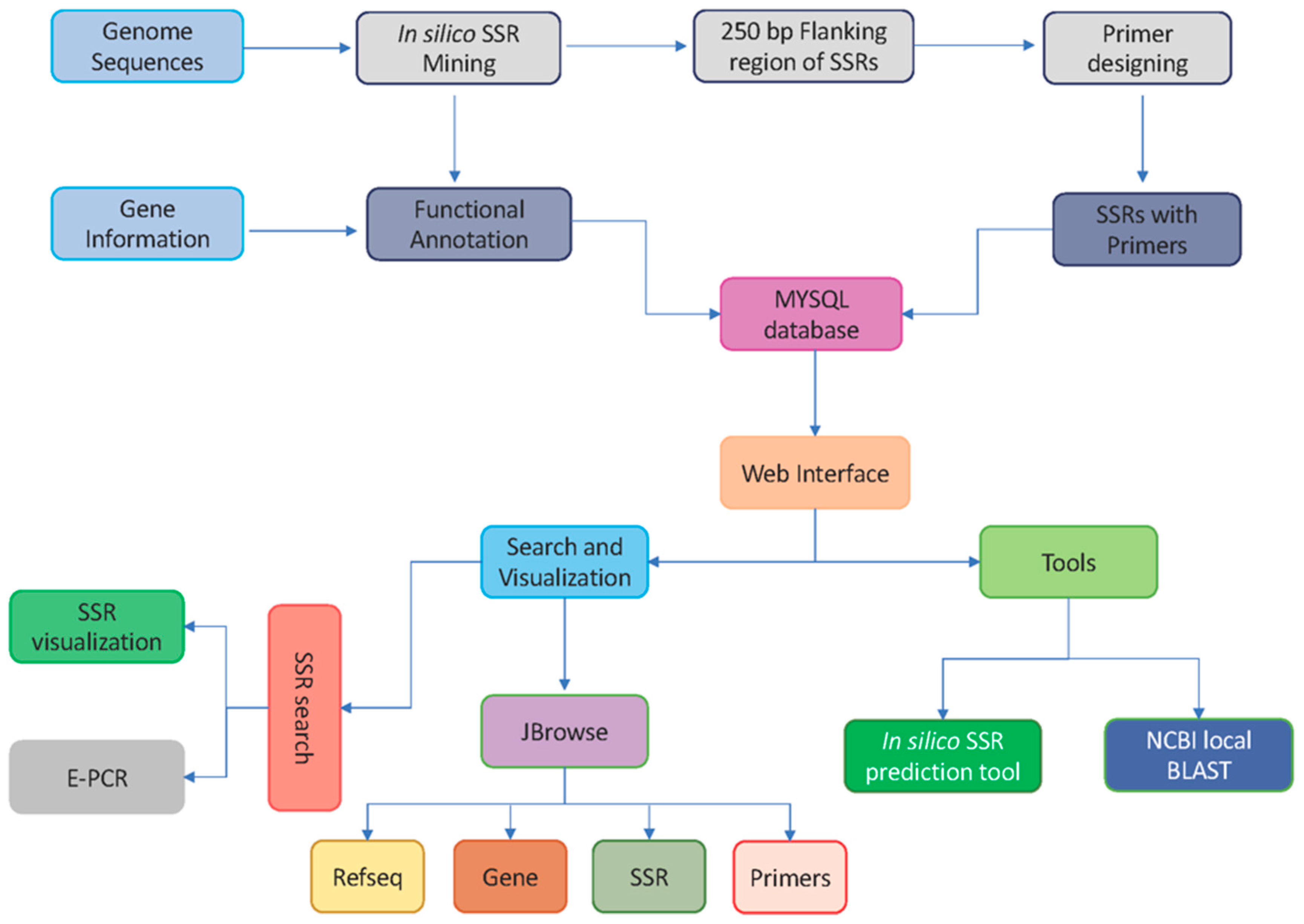
Genes | Free Full-Text | citSATdb: Genome-Wide Simple Sequence Repeat (SSR) Marker Database of Citrus Species for Germplasm Characterization and Crop Improvement

Molecules | Free Full-Text | Mining and Development of Novel SSR Markers Using Next Generation Sequencing (NGS) Data in Plants
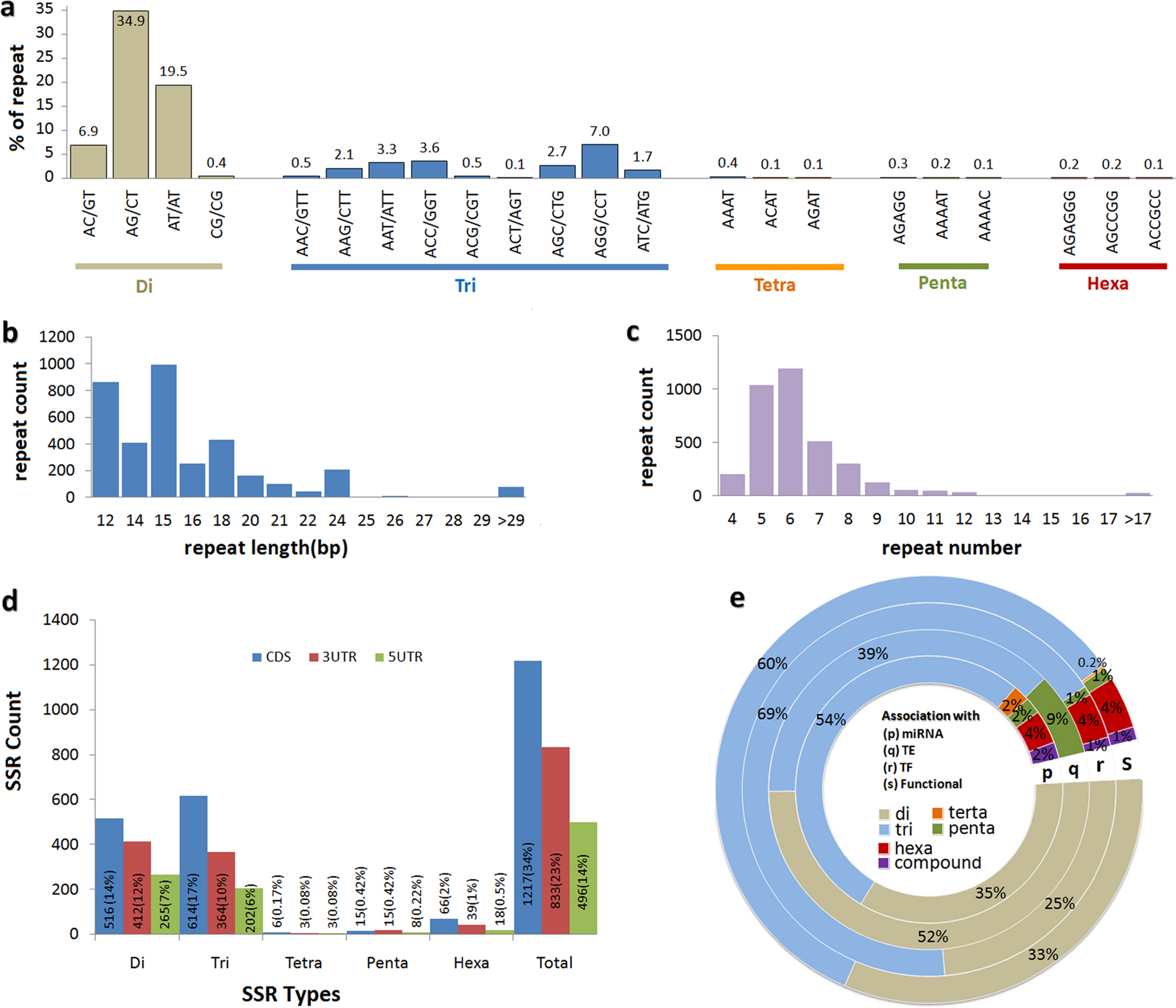
Transcriptome wide SSR discovery cross-taxa transferability and development of marker database for studying genetic diversity population structure of Lilium species | Scientific Reports

Genes | Free Full-Text | Transferability and Polymorphism of SSR Markers Located in Flavonoid Pathway Genes in Fragaria and Rubus Species
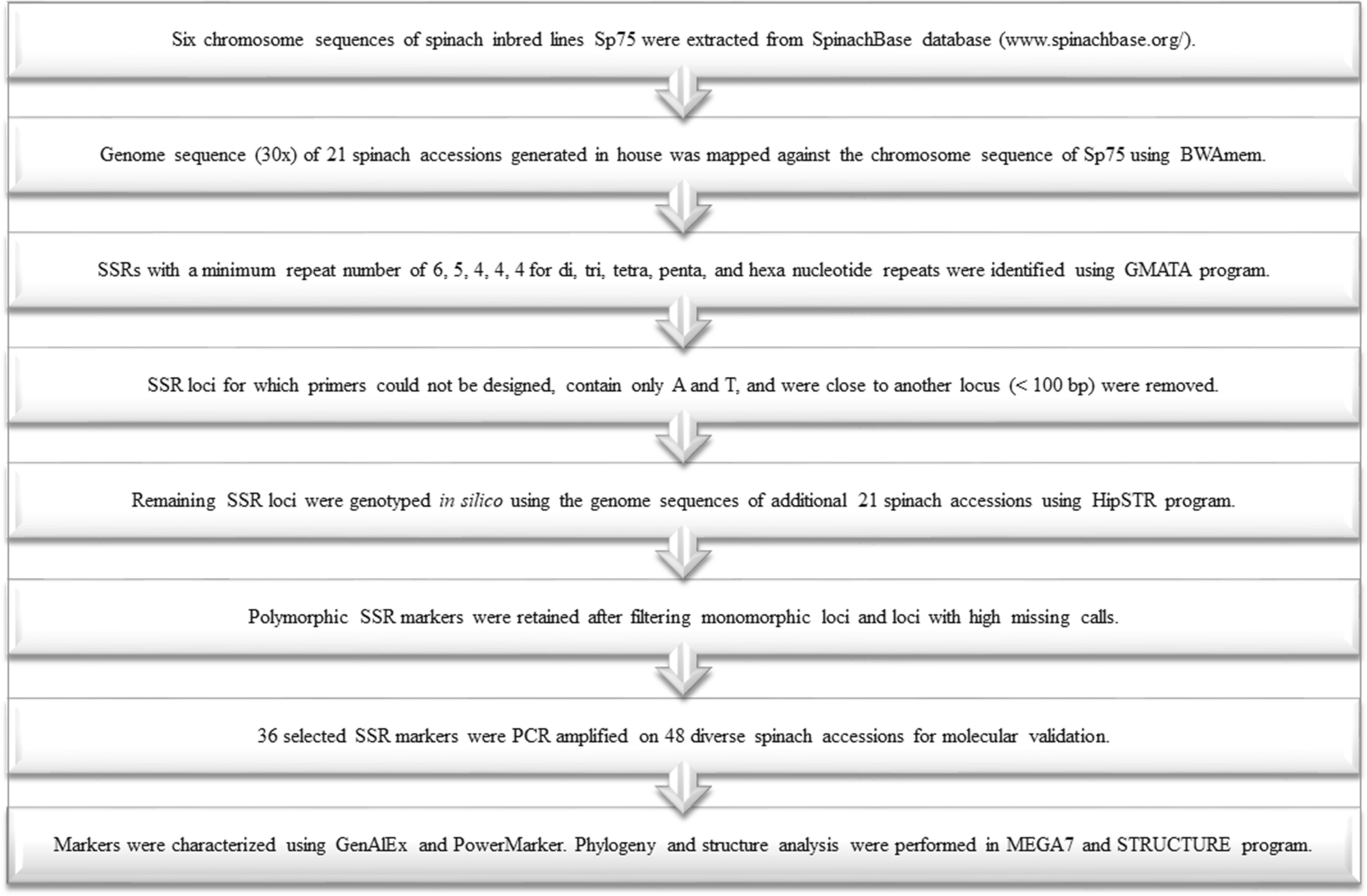
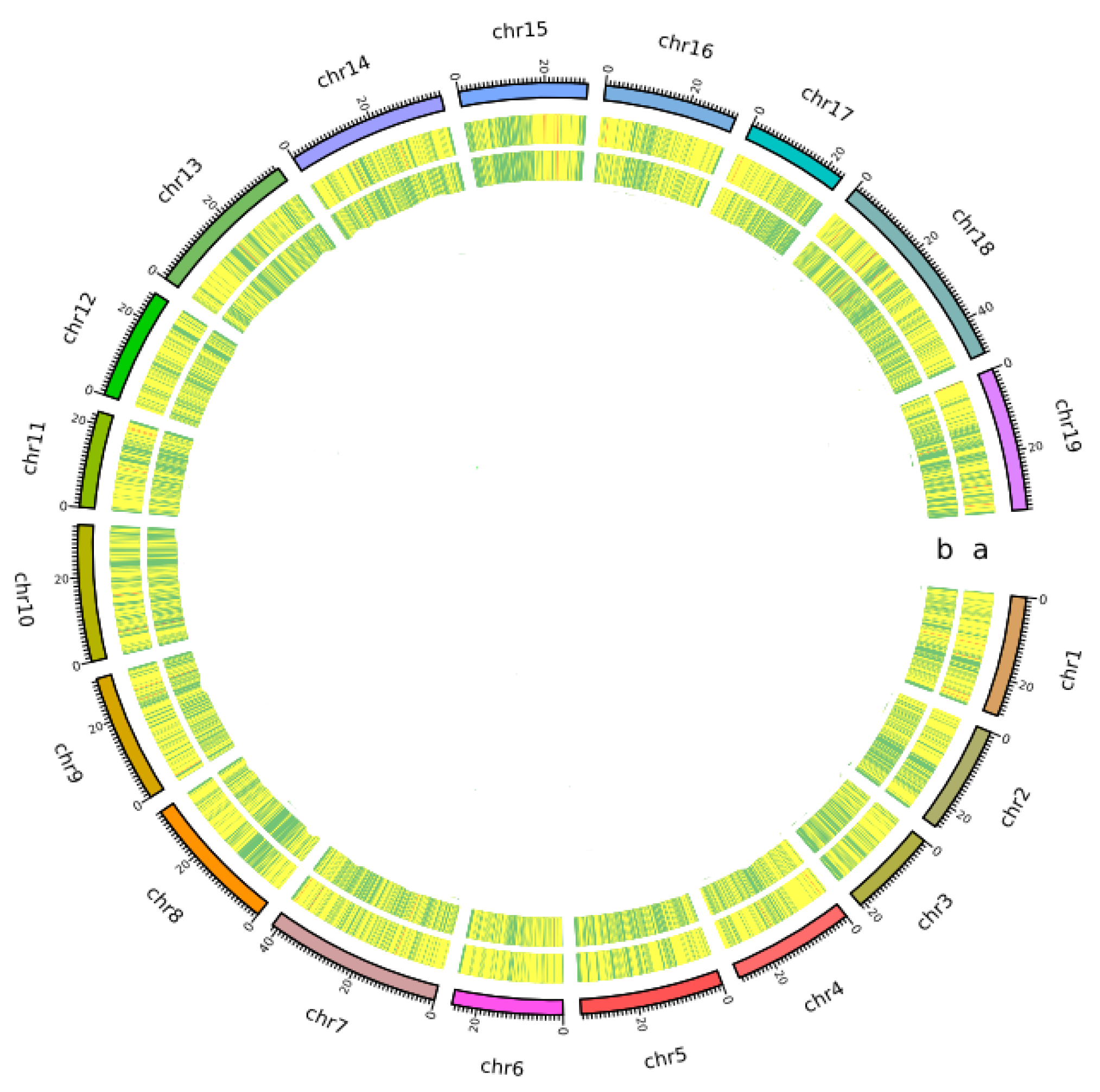
Genome-wide simple sequence repeats (SSR) markers discovered from whole-genome sequence comparisons of multiple spinach accessions | Scientific Reports
Development and Characterization of Simple Sequence Repeat (SSR) Markers Based on RNA-Sequencing of Medicago sativa and In silico Mapping onto the M. truncatula Genome | PLOS ONE

Mining and validation of novel genotyping-by-sequencing (GBS)-based simple sequence repeats (SSRs) and their application for the estimation of the genetic diversity and population structure of coconuts (Cocos nucifera L.) in Thailand

Cross-species transferability of genomic SSR markers and genetic diversity among Asparagus racemosus Willd. Accessions - ScienceDirect
Genes | Free Full-Text | Characterization of Simple Sequence Repeat (SSR) Markers Mined in Whole Grape Genomes

Genome-wide identification of simple sequence repeats and development of polymorphic SSR markers in swamp eel (Monopterus albus) - Hai-feng Tian, Qiao-mu Hu, Zhong Li, 2021